- Proteins are biological polymers composed of amino acids.
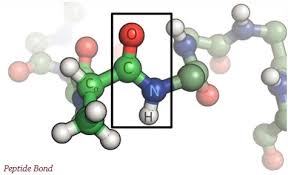
- Amino acids linked together by peptide bond to form a polypeptide chain.
- One or more polypeptide chain twisted into 3D shape to form protein.
- Protein complex shapes includes various folds, loops and curves.
- It is an essential constituent of all cells.
- It is in every part of body, skin, muscles, hair, blood, organs, eyes, finger nails and bones.
- These are macromolecular polymers composed of amino acid as basic unit.

- Protein molecules: fibrous: elongated and insoluble. And Globular: compact, soluble, spherical.
| Globular protein | Fibrous protein |
| Polypeptide chains are compactly folded to form spherical or globular shape | Polypeptide chains are extended along one axis and are spherically wound to form fibers |
| 4 types of bonds: H-bonds, ionic bonds, disulphide bonds and hydrophobic bonds, which maintains a tertiary structure. | H-bonds between amino acids residue, so usually have high degree of secondary structure. |
| Soluble in water | Insoluble in water |
| Non-contractile | Contractile |
| Ex. Egg albumin, globulins, Hb, all enzymes, etc. | Ex. Alpha-keratin of hair, nails, claws, horns, etc. Elastin, Collagen |
| Long, parallel chains form fibres | |

Types of protein:
Simple protein:
Gives only amino acid on hydrolysis. Ex. Ribonuclease
Composed of only alpha-amino acids.
- Albumins: soluble in water and dilute in salt solution. It is present in egg white portion and in blood. It is neutral.
- Globulins: insoluble in water but soluble in dilute salt solution. It is present in antibodies in blood serum and as blood fibrinogen. It is neutral.
- Histones: soluble in water and insoluble in dilute ammonium hydroxide. It is basic in nature. Ex. Chromatin.
Conjugated protein:
Gives amino acid and non- amino acids components on hydrolysis.
Conjugated proteins:
| Class | Prosthetic group | Examples |
| Lipoprotein | Lipids | Beta1- lipoprotein of blood, yolk, serum, milk, and cell membranes |
| Glycoprotein | Carbohydrate | Immunoglobulin G |
| Phosphoprotein | Phosphate group | Casein of milk |
| Hemoprotein | Heme (Fe porphyrin) | Haemoglobin |
| Flavoprotein | Flavin nucleotide | Succinate dehydrogenase |
| Metalloprotein | Fe Zn Ca Mo Cu | Ferritin Alcohol dehydrogenase Calmodulin (any Ca-bindingg protein) Dinitrogenase Plastocyanin |
- Prosthetic group: an inorganic/organic component (non amino acid) that is covalently bound to a protein and essential for its activity.
- Co–factor: an inorganic/organic component is not covalently bound to a protein.
Functional diversity of proteins:
- Enzymes: Hexokinase, kinase, etc.
- Transport proteins: haemoglobin, lipoprotein, membrane transport proteins.
- Nutrient and storage proteins: albumin, casein.
- Contractile protein: actin, myosin, tubulin, etc.
- Structural protein: collagen, desmosine, elastin, keratin, spider web protein.
- Defense protein: immunoglobulin, fibrinogen, thrombin, snake venom, bacterial toxins, rcin, abrin, etc.
- Regulatory protein: hormones, GTP – binding protein.
- Other proteins – antifreeze protein, monellin, etc.
Homologous proteins
Group of protein perform same function in all organism but are evolutionary related (structural resemble)
Example: Hb – oxygen transport
Mb – tissues O2 – transport
Cytochrome – C
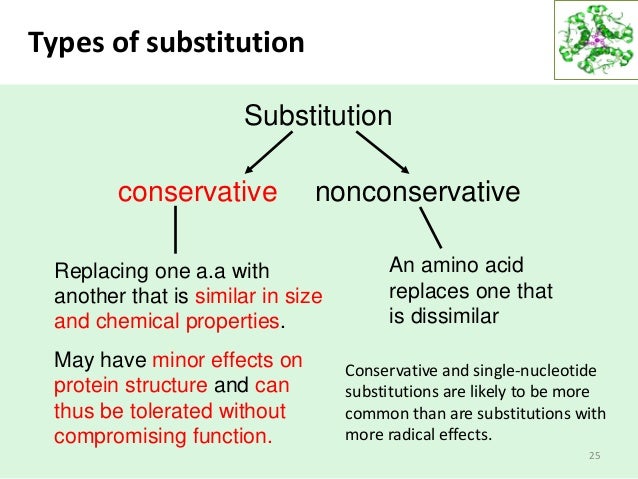
Conservative substitution:
When one amino acid of one part is changed by another. Ex: 1 polar acid changed by 1 another polar amino acid.

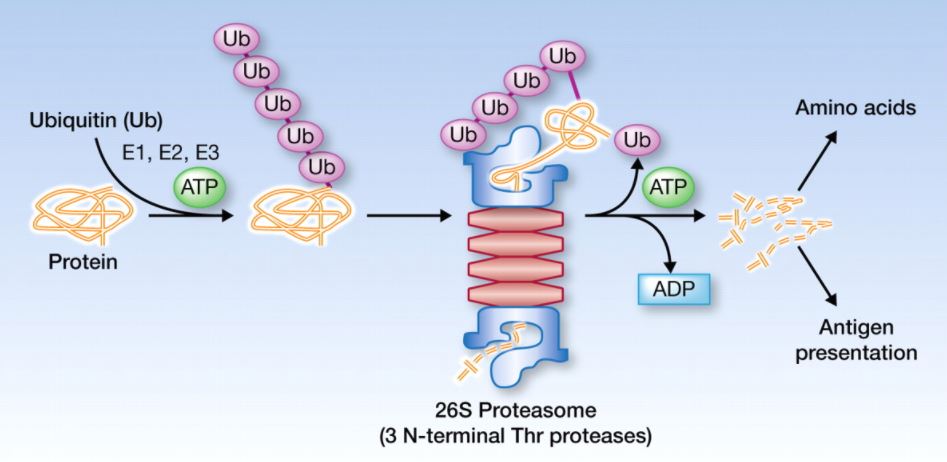
Ubiquitin
With help of ubiquitin, protein degrade after the completion of its function, if ubiquitin do not function then a cell gets block.
Protein conformation
It is stabilized by weak interactions and the stability of unfold state of protein is maintained by high degree of entropy and H-bonding interaction of many group in polypeptide chain.
Conformation:
Spatial arrangement of atoms in a protein and a change in conformation could occur by rotation about single bond.
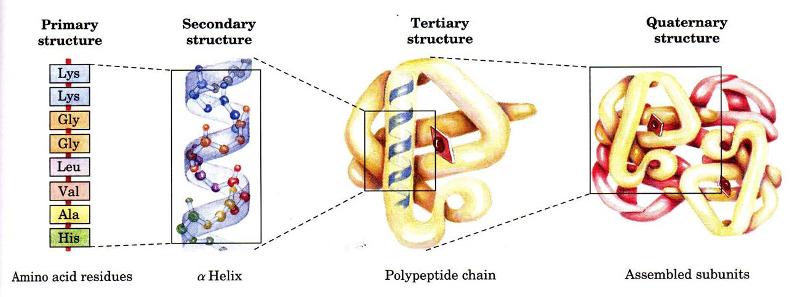
Levels of architecture of proteins:
- Primary structure: it refers to amino acid sequence and all covalent bond between amino acids and location of disulphide bonds.
- Secondary structure: refers to regular recurring arrangement.
- Tertiary structure: refers to spatial relationship between all amino acids.
- Quaternary structure: refers too spatial relationship between different polypeptides.

Primary structure
It is sequence of amino acids in polypeptide chain.
Unique order in which amino acids are linked together to form a protein.
There is the peptide linkage in the form of bonding.
Structure of peptides are: amino acids whose amino group is free and group called N-terminal end, and other whose carboxyl group is free and group called C-terminal end.

Proteins are constructed from a set of 20 amino acids. Generally, amino acids have following structural properties:
C (alpha-C) bonded to 4 group below:-
- H – atom
- -NH2 group
- -COOH group
- Variable or R group
Amino acids sequence of a protein is determined by information found in a cellular genetic code.
Secondary structure
Formed by folding of primary structure.
2 polypeptide are held together by H-bonds.
The coiling or folding of a polypeptide chain that gives proteins its 3D structure.
It is the arrangement od amino acids chain due to the H-bonds between atoms at different parts of the chain.
Ex: collagen and silk – fibrin
Proteins can assume 2 conformational structure: α-helix and β-pleated sheet
α-helix:
- Resembles a coiling spring
- Secured by H-bonding in polypeptide chain.
- Constraints which affect the stability of α-helix:
- Electrostatic repulsions (or attraction) between amino acid residues with charged R-group.
- The bulkiness of adjacent R-group.
- The interaction between amino acid side chain spaced 3 or 4 residues apart.
- Occurrence of proline structure.
- Interaction between amino acid at ends of helix and electric dipole inherent to this structure.
- There is intrachain H-bonding.
Β-pleated sheet:
- Appears to be folded or pleated and is held together by H-bonding between polypeptide units of the folded chain that lie adjacent to one another.
- 2 types:
- Parallel beta sheets:-chains of polypeptides, which run in the same direction.
- Anti-parallel beta sheets:-chains of polypeptides, which run in the opposite direction to eachother.
- These are both intra and inter- chain H bonding, it is extended or zig-zag conformations.

| Α-helix | Β– helix |
| Helical conformation | Zig-Zag conformation |
| Interchain | Inter and Intra- chain |
| R-group project outside | R-group is projected alternately |
Tertiary structure
- Refers to the 3D-structure of polypeptide chains of a protein.
- There are several types of bonds and forces that hold a protein in its tertiary structure.
- Hydrophobic interactions: Greatly contribute to folding and shaping of a protein. ‘R’ group – amino acid: hydrophobic or hydrophilic.
- Hydrophilic R-group: seek contact with aqueous environment
- Hydrophobic R-group: seek to avoid water and position themselves towards the center of protein.
- Hydrogen bonding: In the polypeptide chain and between amino acid R group helps to stabilize protein structure by holding the protein in shape established by hydrophobic interactions.
- Due to protein folding, Ionic bonding can occur between positively and negatively charged R group that come in close contact with one another.
- Folding also results in Covalent bonding between R group of cysteine amino acid. This type of bonding forms: disulfide bridge.
- Interactions called Vander wall forces also assist in stabilization. These contribute to bonding that occurs between molecules.
Quaternary structure
- Proteins are formed by more than one polypeptide chain and joined together by covalent bond.
- Each polypeptide is called as a subunit.
- Quaternary structure formed by interachain.
- When protein formed by similar subunit : homogenous quaternary structure, and when protein formed by dissimilar subunit: heterogenous quaternary structure. It is consists of 2 identical α-chain and 2- identical β-promoter.
- Most stable structure.
Stability
Less stable to more stable:
Primary to secondary to tertiary to quaternary
Chemical bonds involved in protein structure
- Strong bonds
- Peptide bonds: covalent bonds formed by dehydration synthesis between alpha-carboxyl group of one amino acid and alpha-amino group of adjacent amino acid.
- Disulphide bonds: covalent bonds formed between the S-containing cysteine residue of polypeptide chain.
- Weak bonds
- Hydrogen bonds: sharing of H-atoms between the –N and –C=O of different peptide bond.
- Hydrophobic bonds: when two chains of amino acids come together, no true bonds are formed.
- Ionic, electrostatic or salt linkages: formed between closely lying –COO and –NH3 group of different amino group residues.